Jacob Scott Laboratory
-
Jacob Scott Laboratory
- Principal Investigator
- Research
- Our Team
- Publications
- Careers
- Research News

Jacob Scott, MD, DPhil
Staff, Translational Hematology & Oncology Research; Radiation Oncology
Associate Professor, School of Medicine, Case Western Reserve University
Associate Director for Data Science, Case Comprehensive Cancer Center
Adjunct Associate Professor, Department of Physics, Case Western Reserve University
Location: Cleveland Clinic Main Campus
Research
Cancer is a complex disease that is an aberration of our own tissues, but it still obeys fundamental biological rules. Our greatest challenge in the clinic is the emergence of resistance to our therapies, a process which is governed by Darwinian evolution. Using a suite of mathematical and experimental models, my laboratory seeks to deconvolute the complexity of the evolutionary process into fundamental principles. We aim to use this knowledge to then curtail the evolutionary process to increase the efficacy of targeted therapies and radiation. This same knowledge can be further harnessed to understand the differences in disease progression and therapy response on a personalized basis, so that the right treatment can be given to the right patient at the right time.
Biography
I am a veteran of the US Navy submarine force turned academic physician-scientist. Our lab pursues research decomposing the complexity of cancer through mathematical modeling and the biological and clinical validation of these models. My educational background in physics, medicine, mathematics and engineering gives me a unique perspective on cancer and systems biology and I am able to communicate and collaborate with professionals across many disciplines. I have worked extensively on mathematical modeling of cancer evolution and treatment using a variety of models including evolutionary game theory, cellular automata, differential equations and Markov chains. My DPhil thesis focused on the role of heterogeneity, both genetic and microenvironmental, on cancer evolution and radiation response, and my laboratory’s focus is cancer evolution and therapy resistance. Since starting Theory Division, we have begun to diversify, and the lab now has a significant experimental component, conducting experimental evolution in cancer cell lines as well as bacteria. The combination of mathematics, experimental evolution and a clinical focus makes our laboratory stand out as one of the most interdisciplinary in the field of translational cancer evolution and evolutionary medicine. I am eager to use my distinct perspective to help advance this field to help cancer patients.
Education & Professional Highlights
Appointed
2016
Education & Fellowships
Graduate School - St. Anne's College Oxford University
Oxford, OX2, United Kingdom
2018
Residency - H. Lee Moffitt Cancer Center
Radiation Oncology
Tampa, FL USA
2015
Internship - University Hospitals of Cleveland Health System
Internal Medicine
Cleveland, OH USA
2009
Medical Education - Case Western Reserve University
Cleveland, OH USA
2008
Graduate School - Old Dominion University
Engineering Management
Norfolk, VA USA
2003
Undergraduate - United States Naval Academy
Physics
Annapolis, MD USA
1998
Certifications
Radiology - Radiation Oncology
Research
Overview
Theory Division
We now know that cancer proceeds via combined evolutionary and ecological processes. There is irrefutable evidence that populations of cancer cells change as individual cells mutate and selection forces act to shape the population into more fit phenotypes. We now understand that not only do individual cells escape these selection forces by interacting with their environment, including stromal cells like fibroblasts, but also with other tumor cells. However, while basic scientists have established these facts, little, if any, of this knowledge has translated into clinically actionable information; and, more frustratingly, while many scientists have begun to accept these facts, it has not changed the pursuit of ‘silver bullets’: drugs to act on single targets awash in a sea of heterogeneity. In Theory Division, we use a host of tools to approach this problem, with our focus on the evolutionary mechanisms themselves, rather than the specific solutions evolution finds. Below is a snapshot of a few of the things we like to think about... please reach out with questions or ideas - science is a team sport!
Evolution On Adaptive (Fitness) Landscapes
Evolution can be described in many ways. One mathematicall convenient way is consider evolution like a search for peaks (points of highest fitness) on a rugged landscape. If you allow each unique genotype to be a location in space (say the x,y coordinates on a map), then the phenotype is the 'height'. If you now consider each 'map' to be a different drug, you start to reason about how different therapies would affect populations under selection by them. We have modelled bacterial evolution in this way and found that evolution could be effectively 'steered', by a clever ordering of different landscapes, though in reality it is not perfectly controllable because of evolutionary contingencies, but may be predictable. More recently, we have worked to incorporate machinery from statistical mechanics, together with the Hinczewski group in Case Physics to formally derive counter-diabatic driving protocols to allow precise control of the speed and trajectory of populations as they navigate these landscapes.
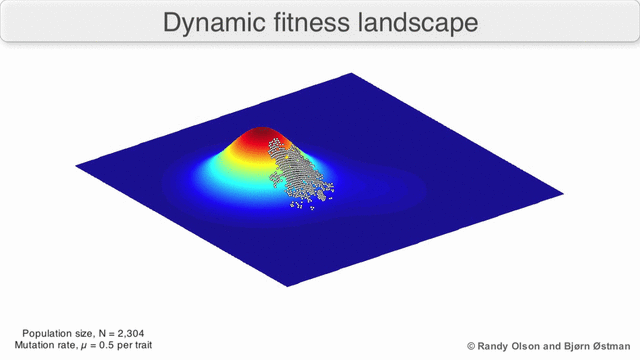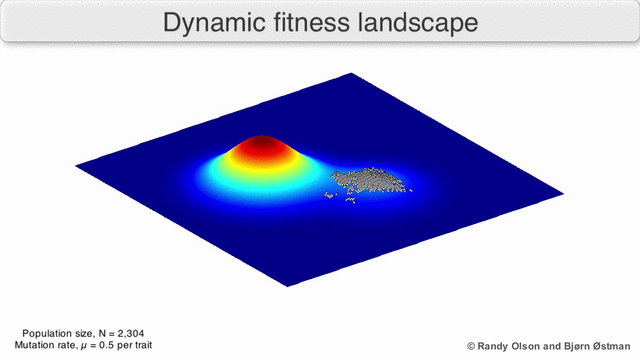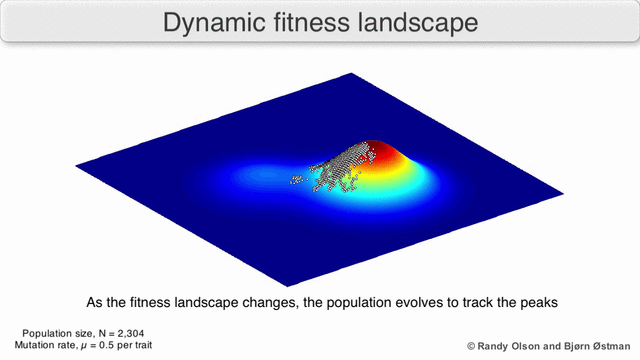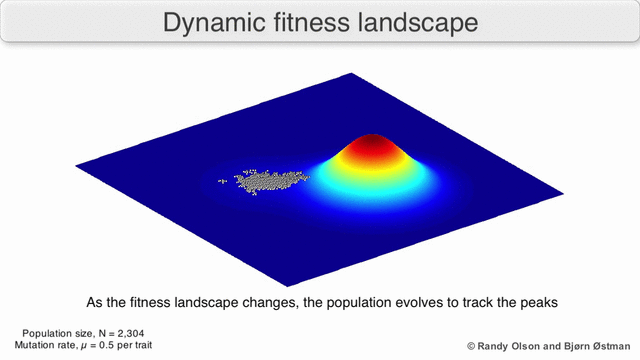
Olson R. and Østman B., "Using fitness landscapes to visualize evolution in action."
Here we show evolution proceeding on three different landscapes in a row. At time 20k and 40k the drugs (landscapes) are switched. In this case, we show an example where the middle drug, cefprozil, is used to steer the evolution of the third drug, ampicillin, to a suboptimal peak. Details can be found here.
Evolutionary Bioreactor
The emergence of drug resistance is the key stumbling block in our fight against cancer. Though our tools can be extremely effective in the short term, it is inevitable that they willfail in the majority of cancers. As an evolutionary phenomenon, we know that the cancer we are treating will “figure out” a way to evade, subvert, and fight back against what we throw at it. We know where the cancer will end up but have relatively little idea how it gets there. We are designing and building a device based on the “morbidostat” framework from the study of antibiotic resistance which we call the EVE system (EVolutionary biorEactor). This will allow us to watch and measure cancer cells on their evolutionary journey towards resistance. It will subject the cells to the same therapies they would experience in patients, but allow us to observe how the cells grow and change much more closely than we ever could in people. The system will be fully robotically automated, providing us a tightly-controlled experimental environment where we can manipulate conditions and see how the cancer cells adjust. Information about the paths that cancers take to resistance can give us new insights about when and how we can intervene while we still can. Our device will allow us to ask and answer questions such as “Can we steer a cancer’s path to slow its progression to resistance?” and “Are there certain times that are better to act than others?”
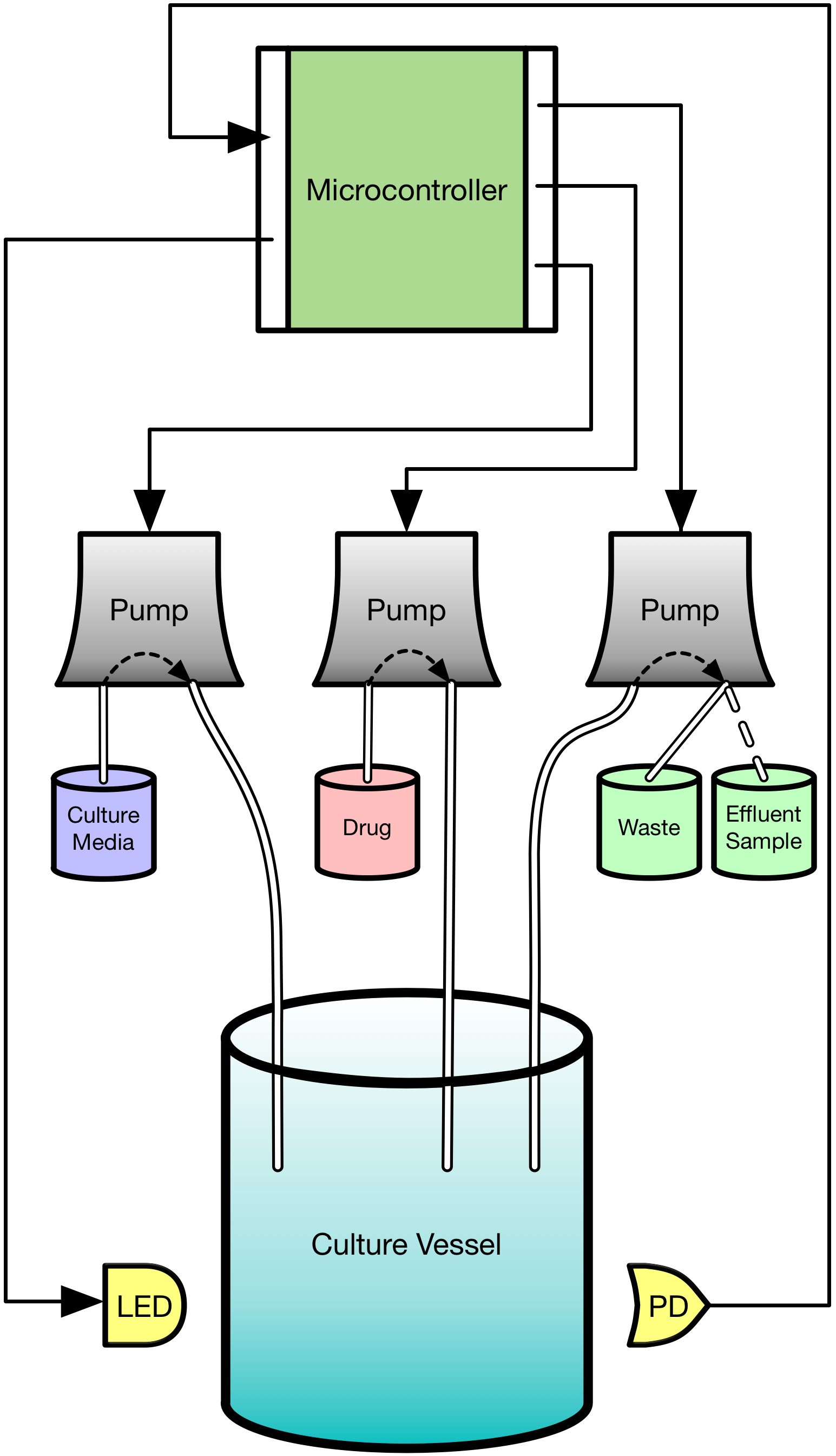
Schematic of evolutionary bioreactor
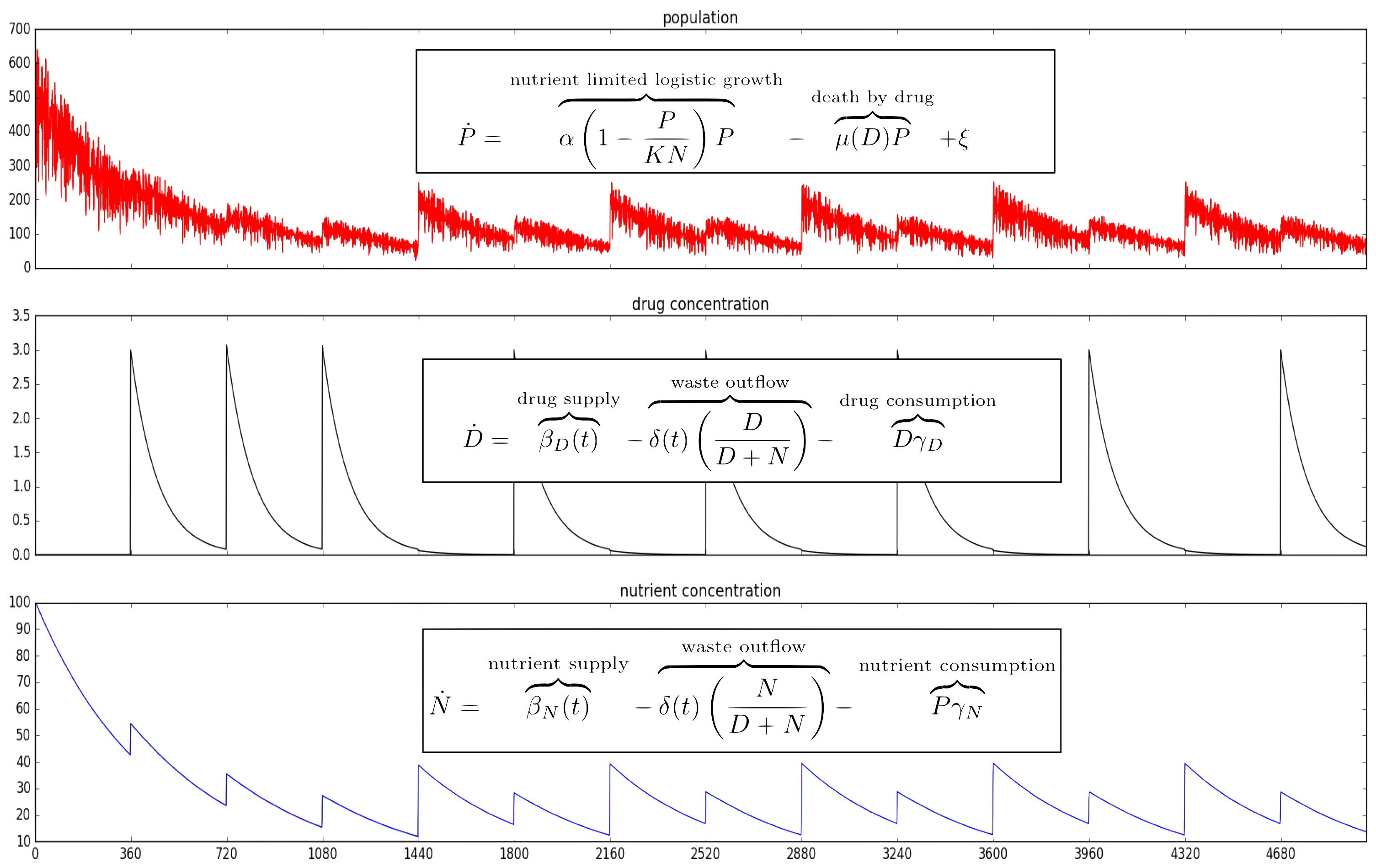
Mathematical model of proposed dynamics
Evolutionary Game Theory
We have an interest in the application of evolutionary game theory to the study of cancer, specifically to the evolution of resistance. Evolutionary game theory allows us to model and perturb the complex cooperative and competitive dynamics between sensitive tumor cells, resistant tumor cells, and the microenvironment. We use mathematical (systems of differential equations) and computational (numerical simulation) models (right), and an experimental game assay we have developed (co-culture and automated cell-counting) (left) to investigate these interactions.
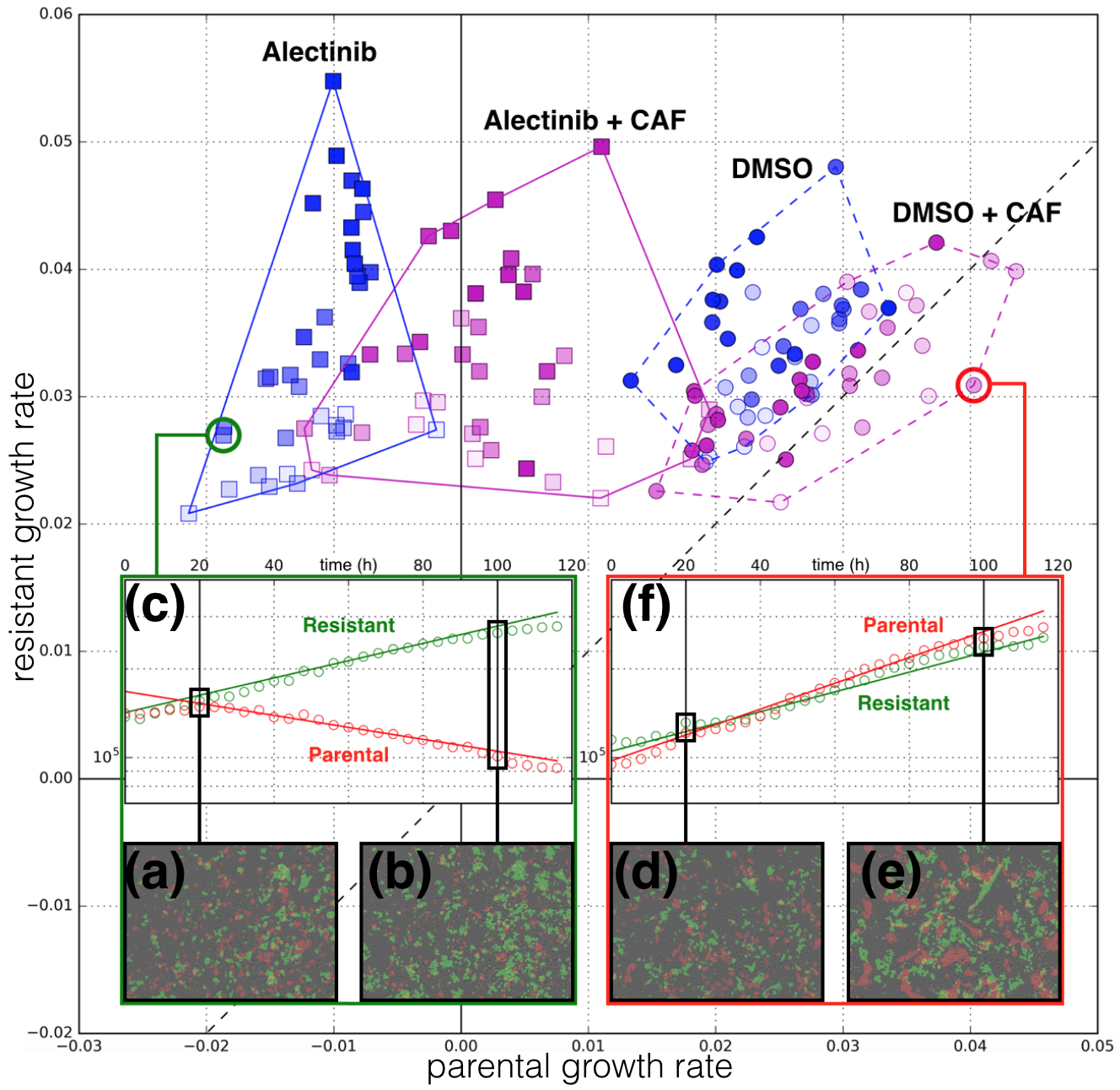
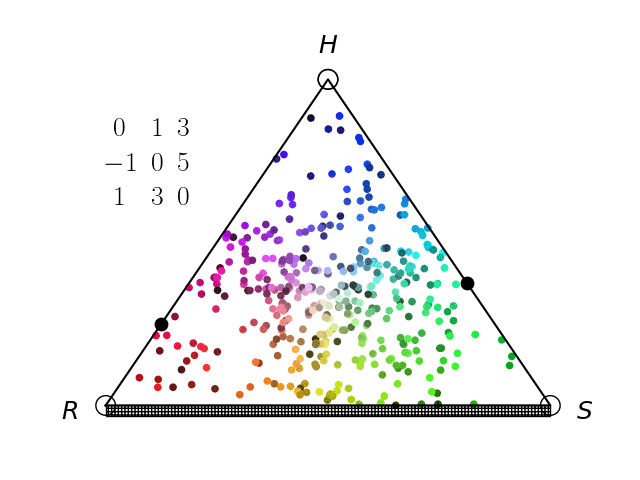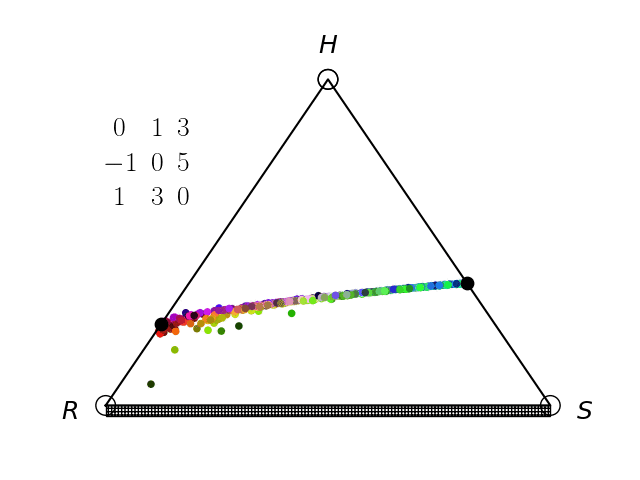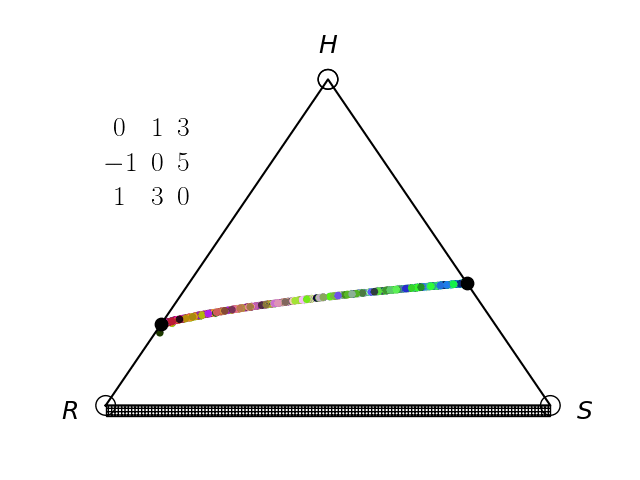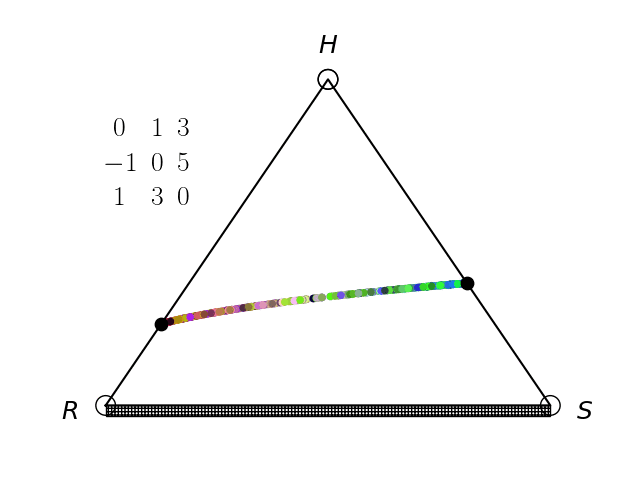
Data Science
With regards to data science, our interests span the breadth of clinical cancer research, ranging from developing novel biomarkers of disease, to selecting personalized therapies (including personalized radiation therapy dosing), and monitoring for resistance. Using datasets encompassing clinico-pathologic, biologic, and genomic data, we hope to design new tests, and lay the groundwork for novel anticancer therapies. To study large high dimensional datasets, we use a combination of high-performance computing and statistical approaches, on problems such as developing gene signatures for drug sensitivity and resistance in Ewings Sarcoma, understanding the functional role of newly characterized non-coding RNA, and determining methods of combining microRNAs into therapeutic cocktails.
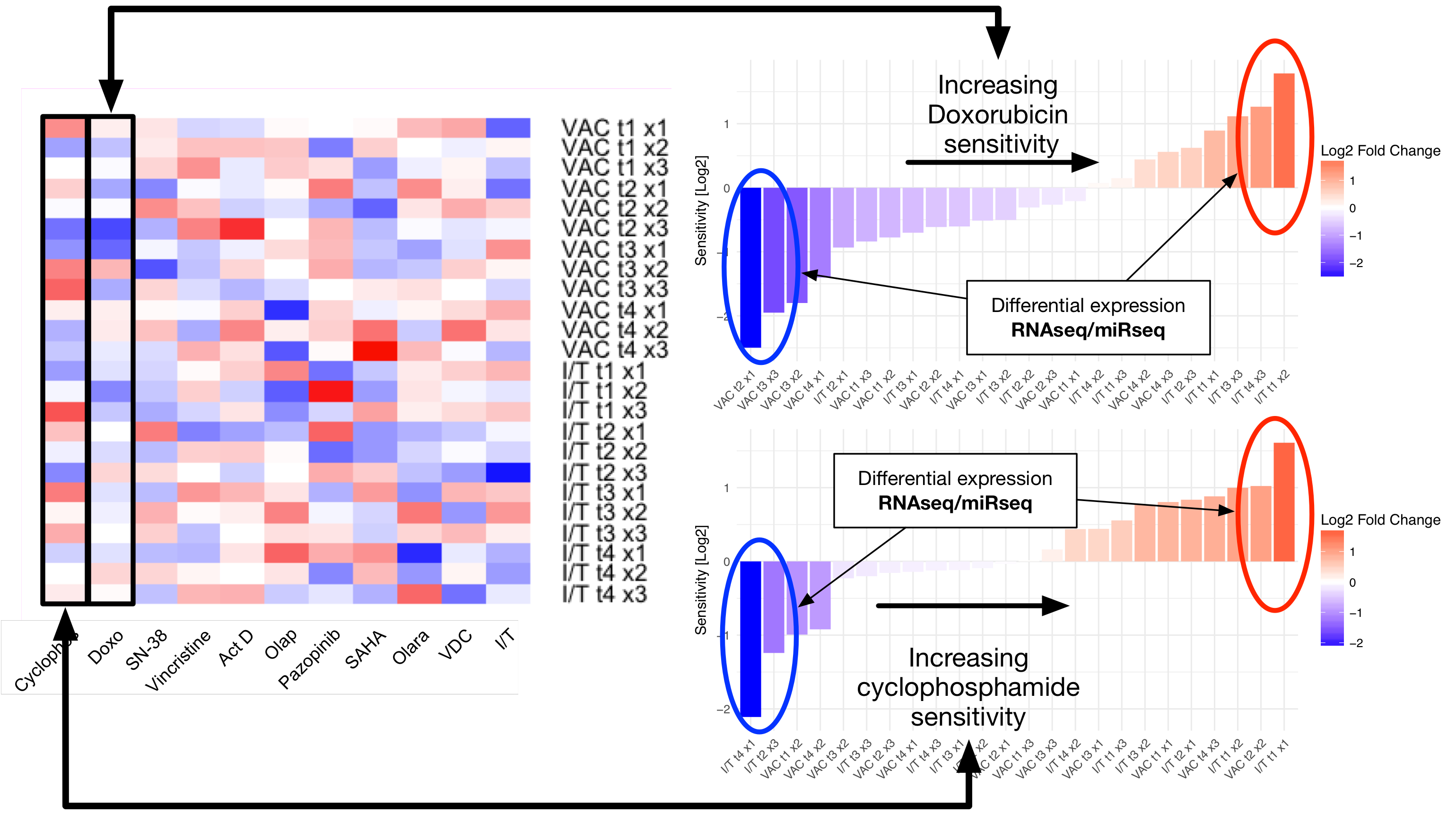
Our Team
Selected Publications
- Weaver Davis, Scott Jacob. Gene Signatures and Oncology Treatment Implications. Hematol Oncol Clin North Am. 2025. 39694780.
- Yang Kailin, Scott Jacob. Unsupervised Bayesian generation of synthetic CT from CBCT using patient-specific score-based prior. Med Phys. 2024. 39666566.
- Dinh Mina N, Hitomi Masahiro, Al-Turaihi Zahraa A, Scott Jacob G. Alamar Blue assay optimization to minimize drug interference and inter assay viability. MethodsX. 2024. 39553738.
- Wei Ruipeng, Hitomi Masahiro, Sadler Tammy, Yehia Lamis, Calvetti Daniela, Scott Jacob, Eng Charis. Quantitative evaluation of DNA damage repair dynamics to elucidate predictors of autism vs. cancer in individuals with germline PTEN variants. PLoS Comput Biol. 2024. 39356721.
- Shultes Paulameena V, Weaver Davis T, Tadele Dagim S, Barker-Clarke Rowan J, Scott Jacob G. Cell-cell fusion in cancer: The next cancer hallmark? Int J Biochem Cell Biol. 2024. 39186970.
- Scott Jacob G. Threshold-awareness in adaptive cancer therapy. PLoS Comput Biol. 2024. 38875286.
- Mayo Zachary S. Meta-Analysis of 5-Fraction Preoperative Radiotherapy for Soft Tissue Sarcoma. Am J Clin Oncol. 2024. 38764405.
- Weaver Davis T, King Eshan S, Maltas Jeff, Scott Jacob G. Reinforcement learning informs optimal treatment strategies to limit antibiotic resistance. Proc Natl Acad Sci U S A. 2024. 38607932.
- Scott Jacob G. Habitat escalated adaptive therapy (HEAT): a phase 2 trial utilizing radiomic habitat-directed and genomic-adjusted radiation dose (GARD) optimization for high-grade soft tissue sarcoma. BMC Cancer. 2024. 38594603.
- King Eshan S, Tadele Dagim S, Pierce Beck, Hinczewski Michael, Scott Jacob G. Diverse mutant selection windows shape spatial heterogeneity in evolving populations. PLoS Comput Biol. 2024. 38386690.
- Kewan Tariq, Durmaz Arda, Bahaj Waled, Gurnari Carmelo, Terkawi Laila, Awada Hussein, Ogbue Olisaemeka D, Ahmed Ramsha, Pagliuca Simona, Kubota Yasuo, Mori Minako, Ponvilawan Ben, Patel Bhumika J, Carraway Hetty E, Scott Jacob, Visconte Valeria, Maciejewski Jaroslaw P. Author Correction: Molecular patterns identify distinct subclasses of myeloid neoplasia. Nat Commun. 2024. 38332139.
- Ågren J Arvid, Scott Jacob G. Viruses, cancers, and evolutionary biology in the clinic: a commentary on Leeks et al. 2023. J Evol Biol. 2023. 37975508.
- Somasundaram Eashwar, Anderson Peter M, Smile Timothy D, Halima Ahmed, Broughman James B, Reddy Chandana A, Scott Jacob G, Chan Timothy, Campbell Shauna, Angelov Lilyana, Zahler Stacey, Trucco Matteo, Thomas Stefanie M, Johnson Shavaughn, Qi Peng, Magnelli Anthony, Murphy Erin S. Neutrophil to lymphocyte ratio (NTLR) predicts local control and overall survival after stereotactic body radiotherapy (SBRT) in metastatic sarcoma. Sci Rep. 2023. 37935813.
- Cho Young-Bin, Suh John H, Scott Jacob G. Radio-immune response modelling for spatially fractionated radiotherapy. Phys Med Biol. 2023. 37459862.
- Somasundaram Eashwar, Wadhwa Raoul R, Barker-Clarke Rowan, Qi Peng, Videtic Gregory, Stephans Kevin, Pennell Nathan A, Raymond Daniel, Yang Kailin, Kattan Michael W, Scott Jacob G. Clinical Nomogram Using Novel Computed Tomography-Based Radiomics Predicts Survival in Patients With Non-Small-Cell Lung Cancer Treated With Stereotactic Body Radiation Therapy. JCO Clin Cancer Inform. 2023. 37369090.
- Smile Timothy D, Halima Ahmed, Broughman James B, Reddy Chandana A, Scott Jacob G, Chan Timothy, Campbell Shauna, Angelov Lilyana, Zahler Stacey, Trucco Matteo, Thomas Stefanie M, Johnson Shavaughn, Qi Peng, Magnelli Anthony, Anderson Peter M, Murphy Erin S. Neutrophil to Lymphocyte Ratio (NTLR) Predicts Local Control Failure and Overall Survival after Stereotactic Body Radiotherapy (SBRT) In Metastatic Sarcoma. Res Sq. 2023. 37333401.
- Scott Jacob G. Towards Data Driven RT Prescription: Integrating Genomics into RT Clinical Practice. Semin Radiat Oncol. 2023. 37331777.
- Kewan Tariq, Durmaz Arda, Bahaj Waled, Gurnari Carmelo, Terkawi Laila, Awada Hussein, Ogbue Olisaemeka D, Ahmed Ramsha, Pagliuca Simona, Kubota Yasuo, Mori Minako, Ponvilawan Ben, Patel Bhumika J, Carraway Hetty E, Scott Jacob, Visconte Valeria, Maciejewski Jaroslaw P. Molecular patterns identify distinct subclasses of myeloid neoplasia. Nat Commun. 2023. 37253784.
- Scarborough Jessica A, Dhawan Andrew, Scott Jacob G. Exploiting convergent phenotypes to derive a pan-cancer cisplatin response gene expression signature. NPJ Precis Oncol. 2023. 37076665.
- Lin-Rahardja Kristi, Weaver Davis T, Scarborough Jessica A, Scott Jacob G. Evolution-Informed Strategies for Combating Drug Resistance in Cancer. Int J Mol Sci. 2023. 37047714.
- Tadele Dagim Shiferaw, Scott Jacob G. Spatial cumulant models enable spatially informed treatment strategies and analysis of local interactions in cancer systems. J Math Biol. 2023. 37017776.
- Mayo Zachary S, Mesko Nathan, Nystrom Lukas, Shah Chirag S, Scott Jacob G, Campbell Shauna R. Radiotherapy innovation in rare diseases-focusing on the value of single institutional experiences for hypofractionated radiotherapy in soft tissue sarcoma. Radiother Oncol. 2023. 36963441.
- Durmaz Arda, Gurnari Carmelo, Hershberger Courtney E, Pagliuca Simona, Daniels Noah, Awada Hussein, Mori Minako, Ponvilawan Ben, Kubota Yasuo, Kewan Tariq, Bahaj Waled S, Barnard John, Scott Jacob, Padgett Richard A, Maciejewski Jaroslaw P, Visconte Valeria. A multimodal analysis of genomic and RNA splicing features in myeloid malignancies. iScience. 2023. 36926651.
- Budd G Thomas, Shepard Dale. Stereotactic Body Radiation Therapy for Sarcoma Pulmonary Metastases. Am J Clin Oncol. 2023. 36914598.
- Huynh Linh, Scott Jacob G, Thomas Peter J. Inferring density-dependent population dynamics mechanisms through rate disambiguation for logistic birth-death processes. J Math Biol. 2023. 36864131.
- Mayo Zachary S, Asha Wafa, Dinh Mina, Mesko Nathan, Nystrom Lukas, Shah Chirag S, Scott Jacob G, Campbell Shauna R. Early outcomes of ultra-hypofractionated preoperative radiation therapy for soft tissue sarcoma followed by immediate surgical resection. Radiother Oncol. 2023. 36481382.
- Gopalakrishnan Vishhvaan, Crozier Dena, Card Kyle J, Krishnan Nikhil P, McClure Erin, Pelesko Julia, Mandal Soumyajit, Bonomo Robert A, Scott Jacob G. A low-cost, open-source evolutionary bioreactor and its educational use. Elife. 2022. 36317871.
- Durmaz Arda, Scott Jacob G. Stability of scRNA-Seq Analysis Workflows is Susceptible to Preprocessing and is Mitigated by Regularized or Supervised Approaches. Evol Bioinform Online. 2022. 36199555.
- Farrokhian Nathan, Maltas Jeff, Dinh Mina, Durmaz Arda, Ellsworth Patrick, Hitomi Masahiro, McClure Erin, Scott Jacob G. Measuring competitive exclusion in non-small cell lung cancer. Sci Adv. 2022. 35776787.
- Yang Kailin, Scott Jacob G. A Phenome-Wide Association Study and the Discovery of a New Clinical Spectrum of Hereditary Cancer Genes. JAMA Oncol. 2022. 35446339.
- Rojas Laura J, Yasmin Mohamad, Marshall Steven M, Krishnan Nikhil P, Perez Federico, Rhoads Daniel D, Jacobs Michael R, Konstan Michael W, Mack Andrew R, Scott Jacob G, Bonomo Robert A. Genomic heterogeneity underlies multidrug resistance in Pseudomonas aeruginosa: A population-level analysis beyond susceptibility testing. PLoS One. 2022. 35358221.
- Sedor Geoff, Scott Jacob G. Changing Radiotherapy Paradigms in Penile Cancer. Eur Urol Open Sci. 2022. 35028598.
- Somasundaram Eashwar, D Smile Timothy, Halima Ahmed, Broughman James B, Reddy Chandana A, Scott Jacob G, Shah Chirag, Chan Timothy, Campbell Shauna, Angelov Lilyana, Anderson Peter M, Zahler Stacy, Trucco Matteo, Thomas Stefanie M, Johnson Shavaughn, Mesko Nathan, Nystrom Lukas, Shepard Dale, Budd George Thomas, Qi Peng, Magnelli Anthony, Murphy Erin S. Association between biologically effective dose and local control after stereotactic body radiotherapy for metastatic sarcoma. J Radiosurg SBRT. 2022. 37416333.
- Anderson Peter M, Thomas Stefanie M, Sartoski Shauna, Scott Jacob G, Sobilo Kaitlin, Bewley Sara. Strategies to Mitigate Chemotherapy and Radiation Toxicities That Affect Eating. Nutrients. 2021. 34959948.
- Scarborough Jessica A, Scott Jacob G. Translation of Precision Medicine Research Into Biomarker-Informed Care in Radiation Oncology. Semin Radiat Oncol. 2022. 34861995.
- Scott Jacob G, Sedor Geoffrey, Scarborough Jessica A, Kattan Michael W. Radiotherapy with genomic-adjusted radiation dose - Authors' reply. Lancet Oncol. 2021. 34735812.
- Weaver Davis T, Scarborough Jessica, Dhawan Andrew, Scott Jacob G. Network potential identifies therapeutic miRNA cocktails in Ewing sarcoma. PLoS Comput Biol. 2021. 34662337.
- Yoon Nara, Krishnan Nikhil, Scott Jacob. Theoretical modeling of collaterally sensitive drug cycles: shaping heterogeneity to allow adaptive therapy. J Math Biol. 2021. 34632539.
- Somasundaram Eashwar, Wadhwa Raoul, Barker-Clarke Rowan, Scott Jacob. Persistent homology of tumor CT scans is associated with survival in lung cancer. Med Phys. 2021. 34587294.
- Scott Jacob G, Sedor Geoffrey, Ellsworth Patrick, Scarborough Jessica A, Kattan Michael W. Pan-cancer prediction of radiotherapy benefit using genomic-adjusted radiation dose (GARD): a cohort-based pooled analysis. Lancet Oncol. 2021. 34363761.
Careers
We are always looking for great folks to work with!
As the primary goal of our lab is to come up with innovative solutions to hard problems, we are interested in scientists with a variety of backgrounds and training. We currently have systems biologists, mathematicians, bioinformaticians, physicists, computer scientists, physicians and a cell biologist.
Graduate Students
I am a certified trainer at Case Western Research University in the Systems Biology and Medical Scientist Training Programs, but accept students from other disciplines as well (math/physics/biology). Interested PhD students should contact Jake with specific questions.
Postdocs
We are always looking for passionate and creative postdocs. We have a slight preference for computationally trained folks, but wouldn't rule out somoene with strong evolutionary or cell biological training who was interested in learning some computation or engineering. If you are interested in working with us, please email Jacob with your CV, a brief description of your background, and your goals in pursuing a postdoc.
Internships, Undergraduates & Visitors
We are always interested in hosting visitors, but time and space are limited, so must choose carefully based on current human capital. Please reach out with questions.
Lab Values & Expectations
Work hard, take care of yourself and others, and be nice.
Lab Culture Statement
Our lab is committed to creating and maintaining an environment in which everyone feels welcome and respected. We value the unique perspectives and experiences each researcher brings and believe they strengthen our team and our work. We are dedicated to removing barriers and ensuring that all members have equal access to opportunities, resources and support.
Mission Statement
The mission of our lab is to create a space for students and postdocs to develop the skills necessary to be able to effectively question, research, and communicate scientific topics. We understand that differences will exist in the level of training, knowledge, personal circumstances, and life and career goals of lab members. As such, our goal is to provide academic opportunity while taking these conditions into account.
Training at Cleveland Clinic Research
Our education and training programs offer hands-on experience at one of the nationʼs top hospitals. Travel, publish in high impact journals and collaborate with investigators to solve real-world biomedical research questions.
Learn MoreResearch News

The researchers demonstrate how vancomycin alters staph evolution, and offer a predictive tool to guide future antibiotic decisions.

Genomic model shows promise in tailoring radiation doses, potentially reducing side effects without compromising cure rates.

Cleveland Clinic researchers are developing a new method of gene expression testing for cancer treatment that tailors drug regimens based on an individual’s chemosensitivity.

Using a form of evolutionary modeling called "seascape models," Cleveland Clinic researchers can predict what happens if you miss a dose of antibiotics.

Understanding how evolution drives resistance to endometrial cancer chemotherapy can help researchers force tumors to evolve vulnerabilities to new treatments.

A new study identifies evolutionary dynamics that support pre-existing treatment resistance mutations in lung cancer and beyond.

Researchers used reinforcement learning to design antibiotic regimens to prevent treatment resistance.

The new Cleveland Clinic-developed model incorporates how evolution contributes to drug effectiveness and dosage.

Work with computer and preclinical models aims to bridge the gap between studying evolutionary biology in the lab and delivering to patient care.

Understanding cell growth dynamics may be the key to controlling therapeutic resistance.